Droplet microfluidics applications
Molecular biology
Droplet microfluidics’ features such as ultra-high rate of producing individual droplets along with its potential for generating nL to pL droplets make it an excellent choice for high throughput molecular biology assays such as:
Single-cell analysis
Population-averaged results in biological assays can be misleading due to the inherent heterogeneity within a population of cells. In cases where the cells of interest are in minority, the majority can mask the target result. It is also conceivable for the response to be dynamic and time-sensitive. These attributions of population-averaged assays call for more advanced tools for single-cell resolution assays.
Undoubtedly, one of the most significant applications of droplet microfluidics devices has been in single-cell analysis. Here, the droplet generator makes droplets comparable to the size of the target cells and encapsulates the cells. Upon encapsulation, the cells can mix with a lysing reagent and undergo one or multiple enzymatic reactions at single-cell resolution. The droplets normally are 10-100 micron in diameter which makes them comparable to the size of eukaryotic cells. Droplet microfluidic chips such as dropseq and indrop amongst the highly popular single-cell analysis microfluidics chips.
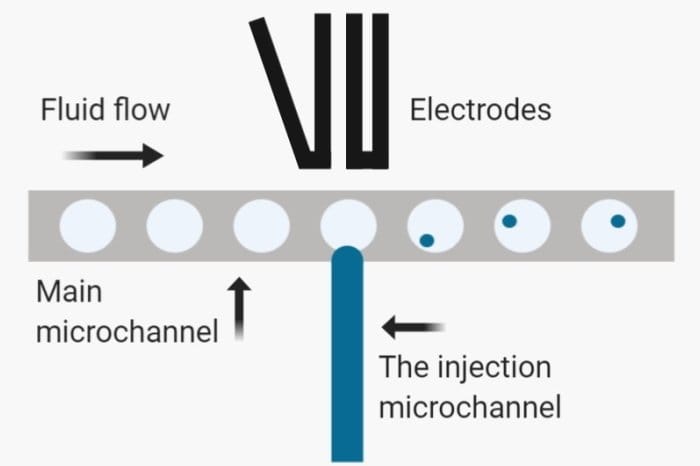
A considerable amount of work in this area has been devoted to expanding this microfluidics technology for the detection and modification of the droplets for single-cell analysis. Fluorescent-based methods for droplet detection are widely used where droplets containing a particular analyte fluoresce and can be distinguished easily. More sophisticated systems incorporate laser data acquisition systems for real-time analysis to detect the fluorescing droplets in real-time. These systems can then be connected to a pair of electrodes to sort the droplets according to the fluorescent signal.
Nucleic acid detection and amplification
The small volume of the droplets makes them an ideal candidate for performing nucleic acid amplification. The droplet confinement acts as the PCR (Polymerase Chain Reaction) chamber and contains the PCR reagents. The droplets will then undergo the thermal cycles necessary for the polymerase chain reaction.
Digital droplet PCR (ddPCR) has been shown to be more accurate than regular qPCR while using the same reagents. Briefly, this microfluidics technique involves segregating the sample into many small pieces followed by a thermal cycling process and statistical analysis of the results. The nature of ddPCR necessitates many parallel assays for high accuracy. Conventional 384 well-plates are common tools in this regard. However, droplet microfluidics, with its capacity for ultra-high droplet generation, has empowered the technique by unprecedented throughput. Thousands of droplets serve as the microchambers for the small pieces of DNA. The result can simply be read out by counting either visually or using image analysis softwares.
Functional material synthesis
 Bulk emulsification procedures had long been the conventional method for producing microspheres. The bulk approaches, however, produce highly polydisperse droplets. Microfluidic technology has attracted a lot of attention for producing functional materials. Droplet microfluidic chips are capable of generating an extremely wide range of droplets that can undergo polymerization to generate microparticles. The sizes vary from several microns to hundreds of microns at ~kHz generation frequencies.
Bulk emulsification procedures had long been the conventional method for producing microspheres. The bulk approaches, however, produce highly polydisperse droplets. Microfluidic technology has attracted a lot of attention for producing functional materials. Droplet microfluidic chips are capable of generating an extremely wide range of droplets that can undergo polymerization to generate microparticles. The sizes vary from several microns to hundreds of microns at ~kHz generation frequencies.
Countless number of hydrogel beads with various sizes have been made with droplet microfluidic technology. The list is not exhaustive but includes PLGA, Agarose, PEG, Alginate, Matrigel, RGD, Collagen, etc.
The microfluidic chips can fabricated to make more advanced particles such as Janus particles (particles with two different materials) and core-shell particles (double emulsions)
Further Reading
- Agarose droplet microfluidics for highly parallel and efficient single molecule emulsion PCR
- A microfluidic chip capable of generating and trapping emulsion droplets for digital loop-mediated isothermal amplification analysis
- High-throughput droplet digital PCR system for absolute quantitation of DNA copy number
- Digital PCR strategies in the development and analysis of molecular biomarkers for personalized medicine
- Microfluidic Generation of Composite Biopolymer Microgels with Tunable Compositions and Mechanical Properties
- Preparation of monodisperse PEG hydrogel microparticles using a microfluidic flow-focusing device
- Microfluidic encapsulation of cells in alginate particles via an improved internal gelation approach
- Janus Microgels Produced from Functional Precursor Polymers
Read more about what needs to be considered before designing a droplet microfluidic chip
There are many microfluidics methods to manipulate droplets after generation. Learn how droplets can be modified.