12 Apr Microfluidic workbench equipped with microtools for on-site enzymatic reactions
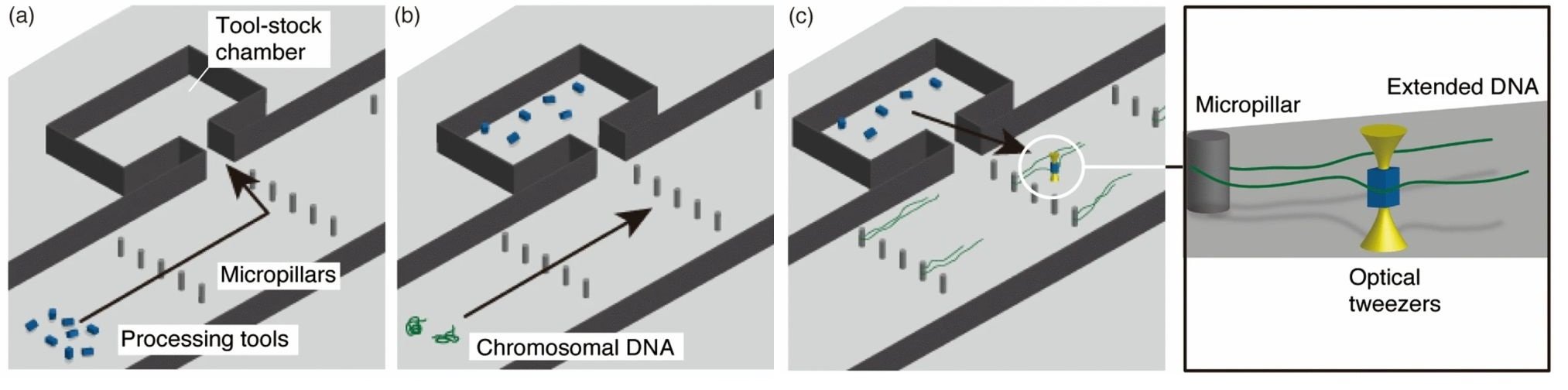
Imagine having a micron-sized operating room equipped with a toolbox full of microtools with which you could perform a micro/nano-operation. A research group in Japan has now taken advantage of microfluidic microfabrication techniques to develop a microfluidic workbench. The microfluidic workbench consists of microchambers equipped with various microtools that can be moved to other reaction chambers to perform a desired set of enzymatic reactions with biomolecules.
“We developed optically driven microtools for processing single biomolecules using a microfluidic workbench composed of a microfluidic platform that functions under an optical microscope. The optically driven microtools have enzymes immobilized on their surfaces, which catalyze chemical reactions for molecular processing in a confined space.”, the authors elaborated.
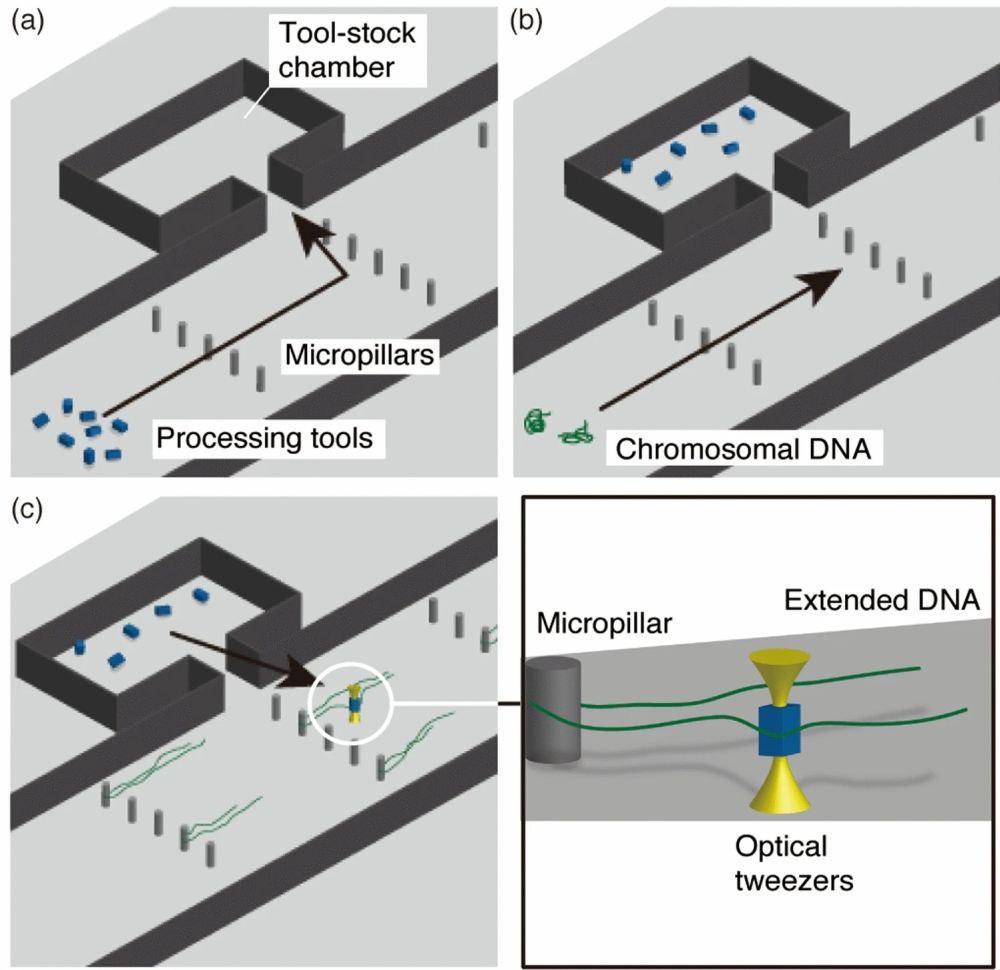
Reproduced under Creative Commons Attribution 4.0 International License. Masuda et al., Sci. Rep., 2021.
The microfluidic workbench was made with PDMS and consisted of a main microchannel beside which side microchambers were placed serving as tool-stock chambers. Micropillars of 6 micron diameter were positioned in the microchannel that helped with trapping and elongating chromosomal DNA flown through the main microchannel. The microtools initially trapped in the side microchambers were brought to the proximity of the chromosomal DNA using optical tweezers to perform the enzymatic reaction.
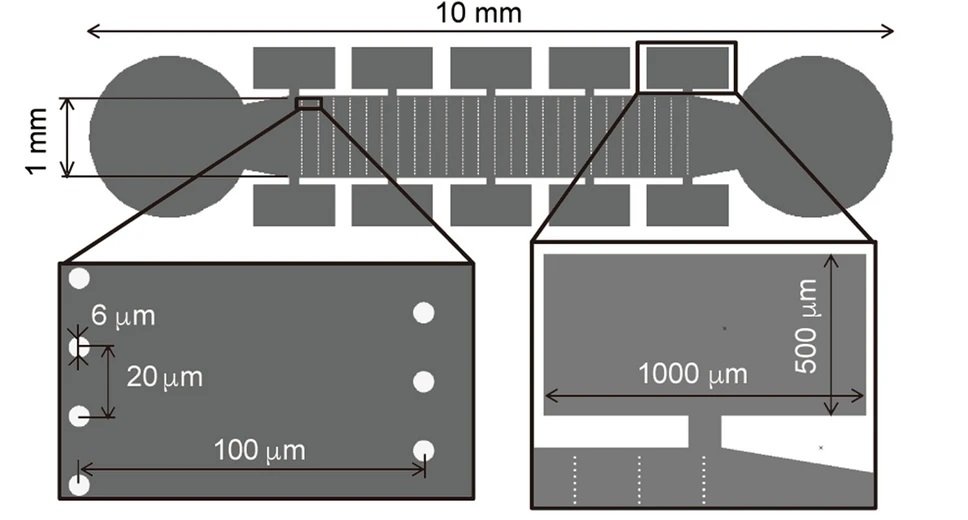
Reproduced under Creative Commons Attribution 4.0 International License. Masuda et al., Sci. Rep., 2021.
The microtools were made off-chip in cuboid shape using SU8. Photolithography was employed to fabricate these cuboids followed by O2/ozone treatment to immobilize biotinylated enzymes. Upon immobilization of the enzymes, the cuboids were scratched from the substrate and collected to be introduced to the side microchambers of the microfluidic device. Two enzymes, DNaseI and DNaseII, were immobilized as a proof of concept. It was shown that the microfluidic chip was capable of cutting the chromosomal DNA. The microtools could be sequentially brought to the reaction microchamber to conduct more complicated enzymatic reactions.
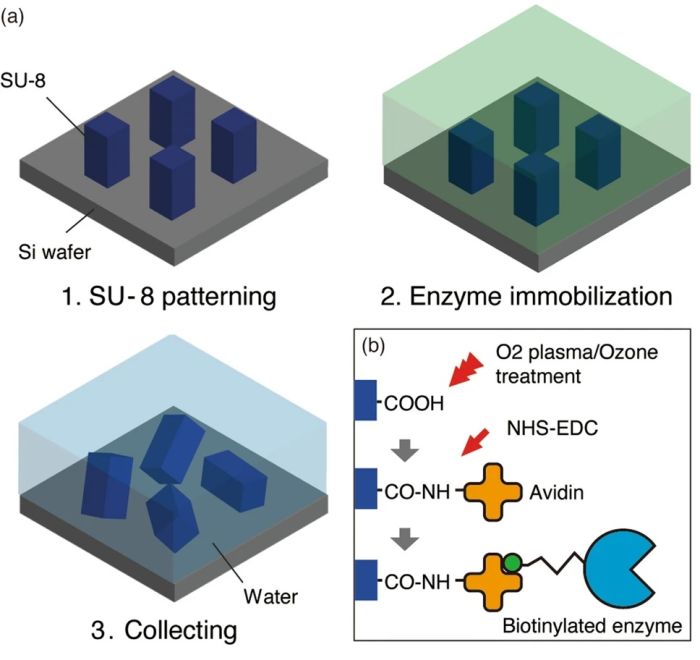
Reproduced under Creative Commons Attribution 4.0 International License. Masuda et al., Sci. Rep., 2021.
“We demonstrated pinpoint chromosomal DNA cutting by DNaseI and DNaseII immobilized microtools, which showed differences in cutting efficiency. The pinpoint processing may provide an approach for analyzing chromosomal DNA at the single-molecule level. The flexibility of the tool and workbench design and surface functionalization will allow the application of a range of pinpoint enzyme reactions to targeted portions of single biosamples. This work provides a basis for the further development of bio-analysis techniques based on single-molecule and single-cell manipulation.”, the authors concluded.
Read the original article: On-site processing of single chromosomal DNA molecules using optically driven microtools on a microfluidic workbench

Pouriya Bayat
Pouriya is a microfluidic production engineer at uFluidix. He received his B.Sc. and M.A.Sc. both in Mechanical Engineering from Isfahan University of Technology and York University, respectively. During his master's studies, he had the chance to learn the foundations of microfluidic technology at ACUTE Lab where he focused on designing microfluidic platforms for cell washing and isolation. Upon graduation, he joined uFluidix to even further enjoy designing, manufacturing, and experimenting with microfluidic chips. In his free time, you might find him reading a psychology/philosophy/fantasy book while refilling his coffee every half an hour. Is there a must-read book in your mind, do not hesitate to hit him up with your to-read list.


