13 Dec Microfluidics and CRISPR in Syncytium Formation: Investigating SARS-CoV-2 Spike Protein Mechanisms
As the world continues to grapple with the challenges of COVID-19, scientists are delving deeper into the virus’s behavior at a cellular level. A pivotal study published in Nature Biomedical Engineering has made significant strides in understanding how the SARS-CoV-2 spike protein induces the formation of syncytia, a process where infected cells fuse to form larger cells. This phenomenon has been a key area of interest in understanding the pathology of the virus. The researchers employed innovative droplet microfluidics alongside CRISPR technology for high-throughput screening. This approach enabled them to identify mutations and cellular determinants that cause the spike protein to induce cell fusion. The integration of microfluidics allowed for the precise handling of minute sample volumes, enhancing the efficiency and accuracy of the screening process.
“Here we describe high-throughput screening methods based on droplet microfluidics and the size-exclusion selection of syncytia, coupled with large-scale mutagenesis and genome-wide knockout screening via clustered regularly interspaced short palindromic repeats (CRISPR), for the large-scale identification of determinants of cell–cell fusion.“, the authors explained.
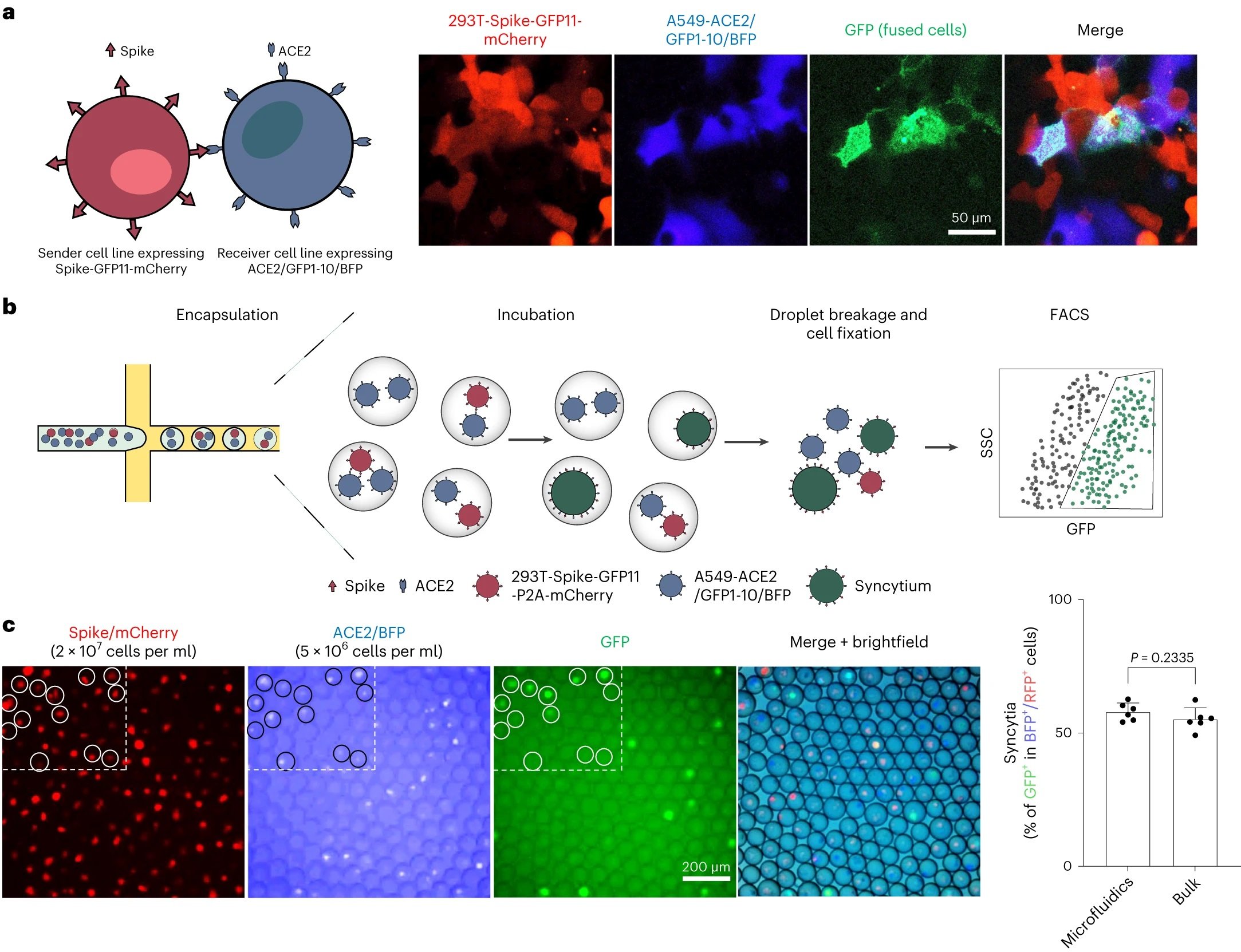
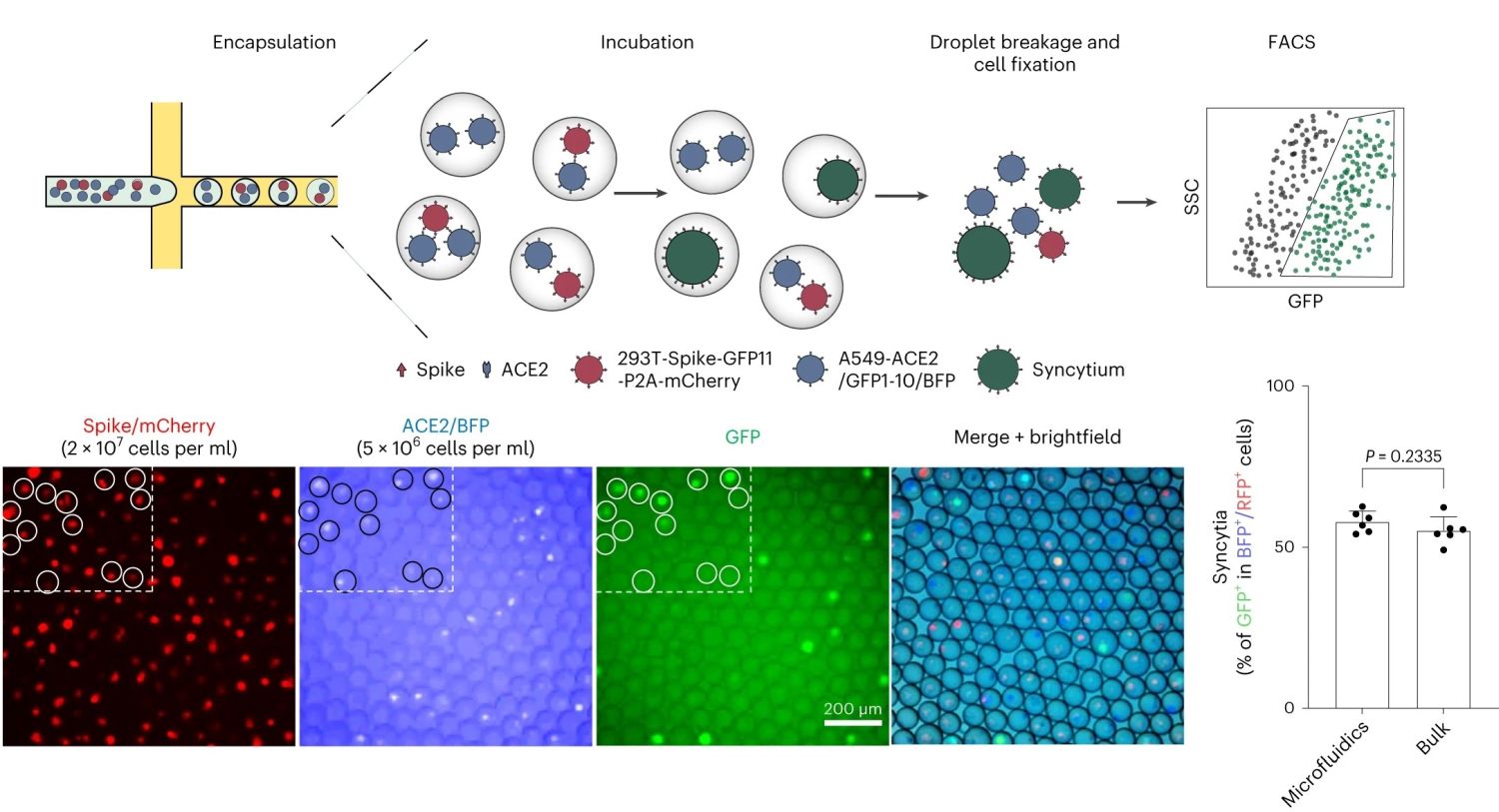
“a, Co-culture of spike-expressing sender cells (mCherry+) and ACE2-expressing receiver cells (BFP+) resulted in syncytium formation (GFP+) due to cell–cell fusion. Scale bar, 50 μm. b, Workflow for cell encapsulation and incubation in droplets and the detection/collection of GFP+ syncytium (boxed). Side scatter (SSC) was used to identify the cell population. c, Representative images of sender cell–receiver cell fusion in droplets. A fusion rate of 57.9% ± 3.3% (data shown are mean ± s.d., n = 6 biological replicates, with P value from unpaired two-tailed Student’s t-test) was measured by FACS after 24 h of incubation in droplets, which is comparable to the bulk setting (55.2% ± 4.3%) in regular co-culture experiments without using droplet microfluidics. Red fluorescent protein (RFP) refers to the mCherry signal detected under FACS.” Reproduced from Chan, C.W.F., Wang, B., Nan, L. et al. High-throughput screening of genetic and cellular drivers of syncytium formation induced by the spike protein of SARS-CoV-2. Nat. Biomed. Eng (2023). under Creative Commons Attribution 4.0 International License.
The study focused on identifying the mutations and cellular determinants that enable the spike protein of SARS-CoV-2 to trigger syncytium formation, a process where infected cells fuse to form larger, multinucleated cells. Syncytia formation is a significant aspect of SARS-CoV-2 pathology and has implications for how the virus spreads and affects the human body. The microfluidic device used in the study was designed for high-throughput screening. Typical microfluidic microfabrication techniques such as soft-lithography was used for microfabricating the microfluidic chips. It incorporated features for droplet creation, allowing for the encapsulation of individual cells or cell clusters within microdroplets. This design facilitated the isolation and detailed analysis of specific cellular behaviors triggered by the virus.
A key feature of the device was its ability to perform size-exclusion selection. This allowed the researchers to segregate syncytia based on their larger size compared to normal cells. The device’s precise control over fluid flow and droplet formation was critical in sorting and analyzing these larger cell structures. Using CRISPR technology, the team conducted genome-wide knockout screenings to pinpoint specific genetic elements influencing cell fusion. They also performed large-scale mutagenesis to explore how various mutations in the spike protein affected syncytium formation. The study carried out deep mutational scans, focusing on two regions of the spike protein: the fusion peptide proximal region and the furin-cleavage site. These scans revealed particular mutations that enhanced the virus’s ability to induce syncytium formation.
The research identified inhibitors of clathrin-mediated endocytosis as key factors that impede syncytium formation. This discovery was validated in animal models, providing potential targets for therapeutic intervention. The findings offer significant insights into the molecular mechanisms of SARS-CoV-2, especially regarding how the virus manipulates cellular processes to spread and exert pathological effects. Understanding these mechanisms is crucial for developing effective treatments. This research exemplifies the power of combining microfluidics with CRISPR technology in viral research. The innovative approach provided a comprehensive understanding of the interactions between SARS-CoV-2 and host cells, paving the way for new strategies to combat COVID-19.
“More broadly, our paired-cell profiling systems can be applied to study a variety of pathological, physiological and even synthetic conditions that are relevant to biomedical applications. Apart from SARS-CoV-2, a broad spectrum of viruses including human immunodeficiency virus, Herpesviridae, respiratory syncytial virus, as well as other Coronaviridae, induces syncytium formation. “, the authors concluded.
For more insights into the world of microfluidics and its burgeoning applications in biomedical research, stay tuned to our blog and explore the limitless possibilities that this technology unfolds.
Figures are reproduced from Chan, C.W.F., Wang, B., Nan, L. et al. High-throughput screening of genetic and cellular drivers of syncytium formation induced by the spike protein of SARS-CoV-2. Nat. Biomed. Eng (2023). https://doi.org/10.1038/s41551-023-01140-z under a Creative Commons Attribution 4.0 International License)
Read the original article: High-throughput screening of genetic and cellular drivers of syncytium formation induced by the spike protein of SARS-CoV-2


