29 Mar One-step RT-PCR for ultrahigh-throughput single-cell analysis using droplet microfluidic chips
Single-cell analysis has become one of the most important applications of microfluidic technology in recent years. It refers to the process of isolation, manipulation, and analysis of the cells at single-cell resolution in microfluidic chips. Depending on the application, single-cell analysis chips can include any or all the above-mentioned steps. The droplet microfluidic single-cell analysis enables researchers to distinguish differences between individual cells as well as detection of rare cells within a seemingly homogenous sample. Although various single-cell analysis techniques have been introduced over the past several years, the cost and sensitivity are still major challenges that researchers face when it comes to the practice.
A multidisciplinary research group has now developed a microfluidic platform to make single-cell analysis more accessible. They took advantage of the high-throughput droplet-based microfluidic and combined it with a single-cell RT-PCR platform to develop a low-cost device capable of analyzing multiple targets in single-cell samples from a large population in one step.
“Here, we present a microdroplet-based, multiplex RT-PCR platform that is designed for ultrahigh-throughput single-cell analysis of differential gene expression as well as rare cell populations. This technique provides single-cell resolution information with high signal-to-noise ratio and acquisition of hundreds of thousands of data points, which allows the analysis of rare cell populations down to a frequency of at least 0.1%.”, the authors elaborated.
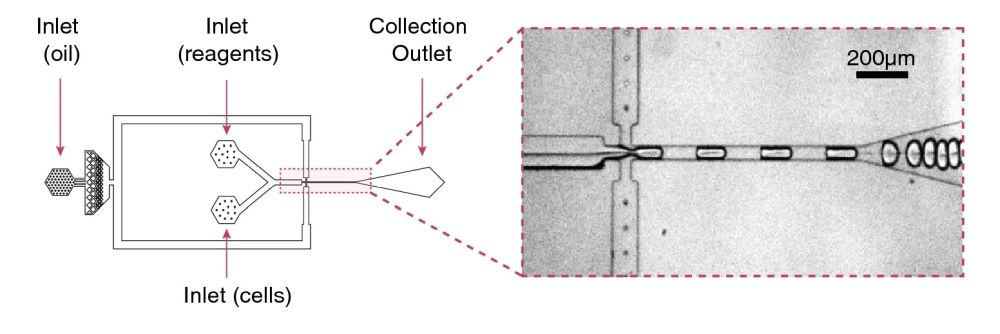
Reproduced under Creative Commons Attribution 4.0 International License. Ma et al., Sci. Rep., 2021.
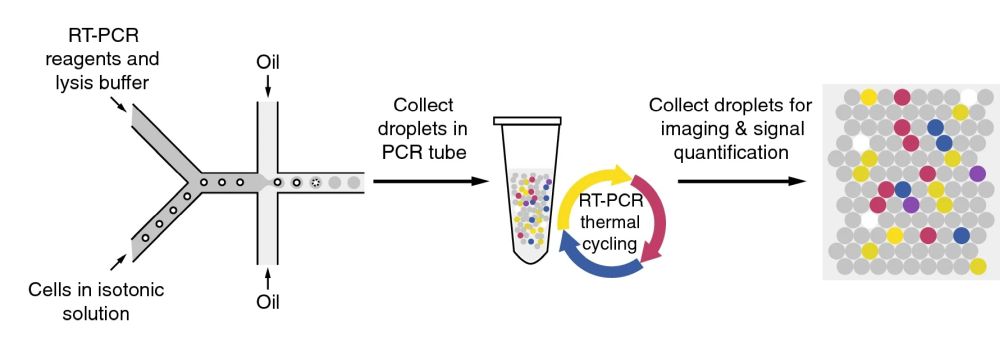
A PDMS-based microfluidic device was microfabricated using standard and photolithography methods for droplet generation. The microfluidic chip generated nL droplets in which single cells were encapsulated along with lysis buffer and RT-PCR reagents. The microfluidic chip had two separate inlet ports to deliver the cells and the reagents to the chip separately. By employing a microfluidic droplet generator, single cells can be encapsulated at extremely high rates. In this case, 1 million droplets of ~124 micron diameter were generated in an hour. Before forming the emulsion, the cell suspension and the reagents had the chance to mix. High concentrations of cell lysates can inhibit the amplification. This issue was resolved by employing an in-house reagent mix containing bacteriophage T7 gene 2.5 protein (gp2.5). The droplets were then collected in a PCR tube and underwent thermal cycling. The droplets were then pipetted into microchambers made from PMMA sheets to form monolayers for fluorescent imaging.
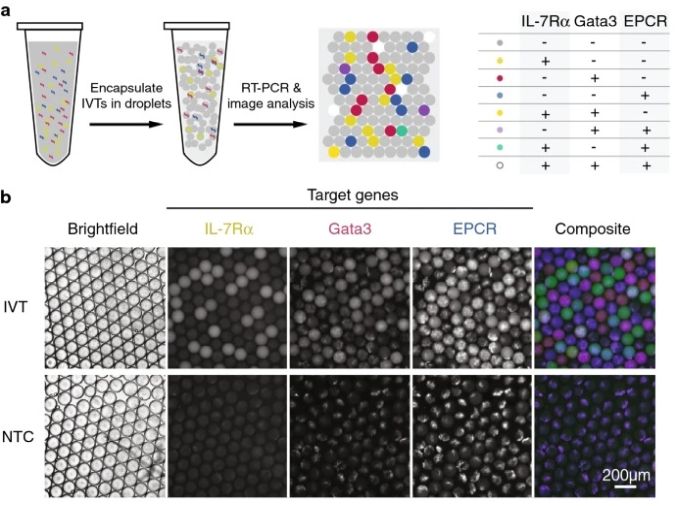
Reproduced under Creative Commons Attribution 4.0 International License. Ma et al., Sci. Rep., 2021.
There are various means for errors to be introduced to the platform. For example, ambient RNA could get co-encapsulated in the droplets and result in a false positive. Additionally, cells encapsulated using droplet microfluidic chips follow a Poisson distribution. The Poisson distribution determines the probability of having droplets containing a certain number of cells and is dependant on the concentration of the cells as well as the droplet volume. Based on the Poisson distribution, there is always a possibility -although very low- to have droplets with more than one cell which can cause errors in the final analysis of gene structure profiles. These errors were corrected by incorporating a deconvolution pipeline that corrected for ambient RNA and multiplets. The microfluidic platform was shown to be capable of one-step RT-PCR of multiple targets at almost $3 per 1000 cells which makes it an affordable test that can be conducted in standard biology labs.
“In summary, our unique platform takes advantage of the miniaturization, compartmentalization, and precise liquid handling capabilities of the ultrahigh-throughput microdroplet system to develop an automatable, low-cost, and highly sensitive assay. As such, this novel technique provides a better single-cell analysis solution for many applications than other existing platforms, and has a high potential to be implemented in both clinical and research settings for rapid and direct evaluation of single-cell gene expression.”, the authors concluded.
Read the original article: Microdroplet-based one-step RT-PCR for ultrahigh throughput single-cell multiplex gene expression analysis and rare cell detection

Pouriya Bayat
Pouriya is a microfluidic production engineer at uFluidix. He received his B.Sc. and M.A.Sc. both in Mechanical Engineering from Isfahan University of Technology and York University, respectively. During his master's studies, he had the chance to learn the foundations of microfluidic technology at ACUTE Lab where he focused on designing microfluidic platforms for cell washing and isolation. Upon graduation, he joined uFluidix to even further enjoy designing, manufacturing, and experimenting with microfluidic chips. In his free time, you might find him reading a psychology/philosophy/fantasy book while refilling his coffee every half an hour. Is there a must-read book in your mind, do not hesitate to hit him up with your to-read list.


