30 Jun Droplet microfluidics-enabled total RNA sequencing
Abstract
“Most methods for single-cell transcriptome sequencing amplify the termini of polyadenylated transcripts, capturing only a small fraction of the total cellular transcriptome. This precludes the detection of many long non-coding, short non-coding and non-polyadenylated protein-coding transcripts and hinders alternative splicing analysis. We, therefore, developed VASA-seq to detect the total transcriptome in single cells, which is enabled by fragmenting and tailing all RNA molecules subsequent to cell lysis. The method is compatible with both plate-based formats and droplet microfluidics. We applied VASA-seq to more than 30,000 single cells in the developing mouse embryo during gastrulation and early organogenesis. Analyzing the dynamics of the total single-cell transcriptome, we discovered cell type markers, many based on non-coding RNA, and performed in vivo cell cycle analysis via detection of non-polyadenylated histone genes. RNA velocity characterization was improved, accurately retracing blood maturation trajectories. Moreover, our VASA-seq data provide a comprehensive analysis of alternative splicing during mammalian development, which highlighted substantial rearrangements during blood development and heart morphogenesis.”
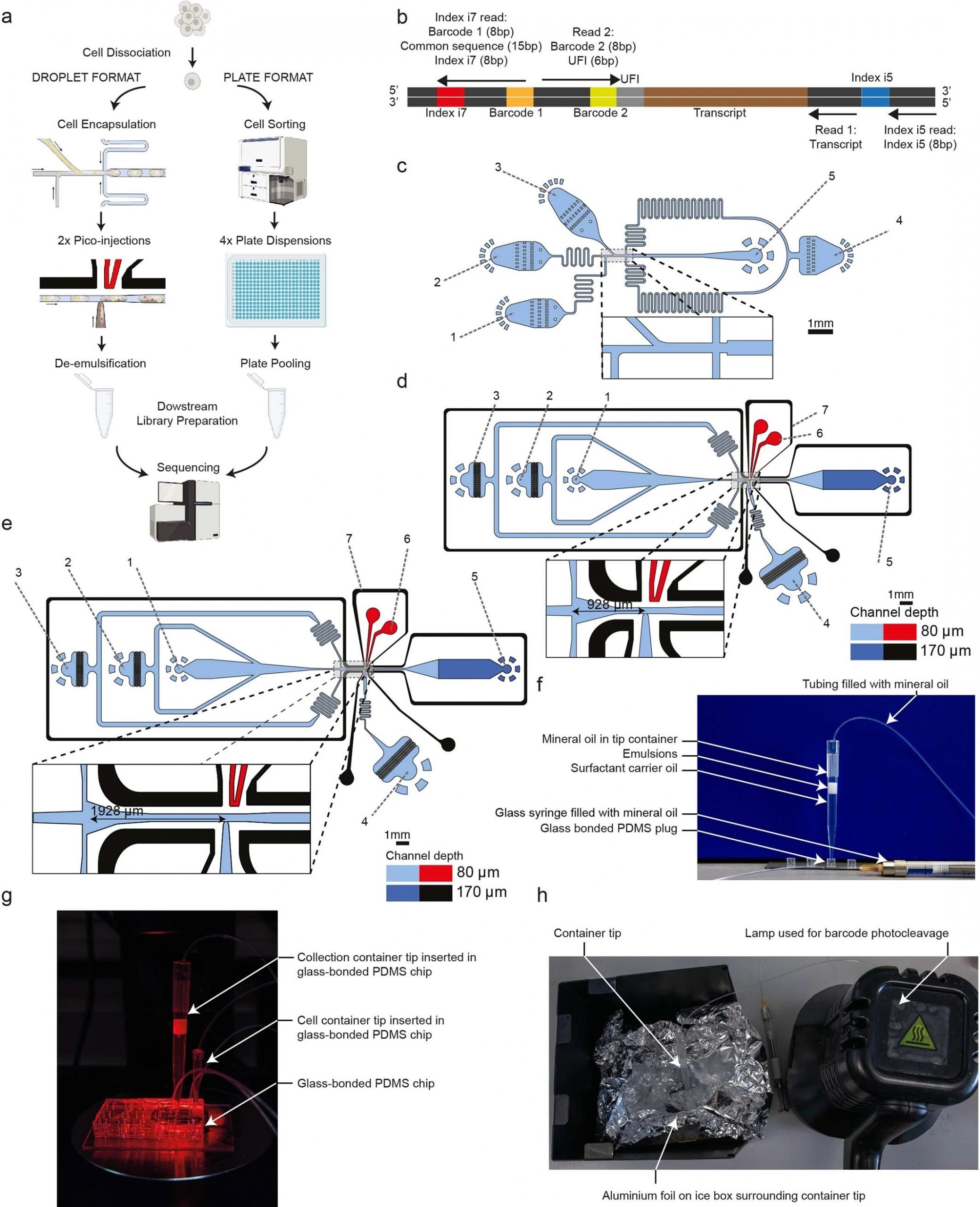
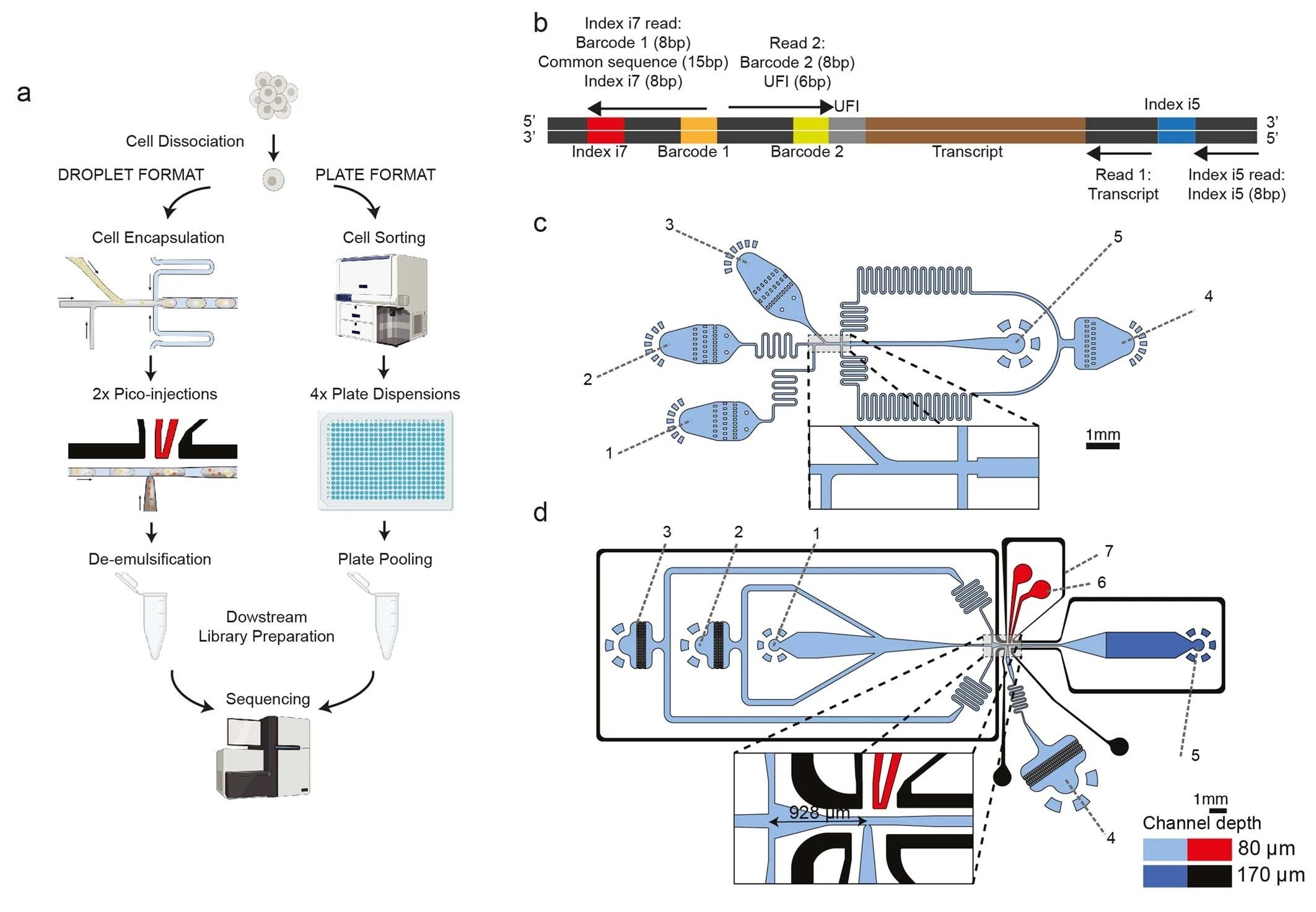
“a, The two platforms for VASA-seq, using a microfluidic device (left, VASA-drop) and a plate dispenser (right, VASA-plate). The microfluidic device allows the generation of single-cell libraries from thousands of cells, while the plate-based approach is better for rare cell types where a prior sorting is required. Each library contains the transcriptome from a large mix of cells, which are demultiplexed based on their barcode and index sequences. b, Library barcoding design for the VASA-drop workflow. Mouse embryo libraries were sequenced with the Illumina NovaSeq platform. To avoid index hopping, a custom dual indexing strategy was used. For the index i7 read, which usually only contains barcode 1 (inDrop v3), we inserted a 8-bp second index directly after a 15 bp common sequence. Only reads that had the correct combination of i5 and i7 index were further used for downstream processing. c, Design of the device used for barcoded bead and single-cell co-encapsulation. 1) input channel for the lysis and fragmentation mix, 2) input channel for the cell loading, 3) input channel for the barcoded compressible bead loading, 4) input channel for the fluorinated oil with admixed surfactant, 5) droplet exit channel. d, Design of the first picoinjector device, to inject the repair and poly(A) polymerase. 1) input channel for droplet reinjection, 2) emulsion diluting oil inlet, 3) droplet spacing oil inlet, 4) inlet for the repair and poly(A) polymerase to be picoinjected, 5) droplet exit channel, 6) positive electrode (red), 7) negative (moat) electrode (black). e, Design of the second picoinjector device, to inject the RT enzyme mix. 1) input channel for droplet re-injection, 2) emulsion diluting oil inlet, 3) droplet spacing oil inlet, 4) inlet for the RT mix to be picoinjected, 5) droplet exit channel, 6) positive electrode (red), 7) negative (moat) electrode (black). f, Photography of the container tip used for droplet collection and reinjection in the picoinjector devices. The tip is connected to a glass syringe which enables aspiration and delivery of emulsions. The tip can be connected to a PDMS plug to close the system for incubation in the water bath. g, Photography of the encapsulation process. The container tip collecting the emulsions is plugged into the outlet of the encapsulation device, while a tip containing cells is plugged in one of the inputs to deliver single-cells in the droplets. h, Photography of the bead barcode photocleavage set-up. The container tip is surrounded by aluminium foil to reflect the UV light and is kept on a container filled with ice.” Reproduced under Creative Commons Attribution 4.0 International License from Salmen, F., De Jonghe, J., Kaminski, T.S. et al. High-throughput total RNA sequencing in single cells using VASA-seq. Nat Biotechnol (2022).
Figures and the abstract are reproduced from Salmen, F., De Jonghe, J., Kaminski, T.S. et al. High-throughput total RNA sequencing in single cells using VASA-seq. Nat Biotechnol (2022). https://doi.org/10.1038/s41587-022-01361-8 under Creative Commons Attribution 4.0 International License.
Read the original article: High-throughput total RNA sequencing in single cells using VASA-seq


